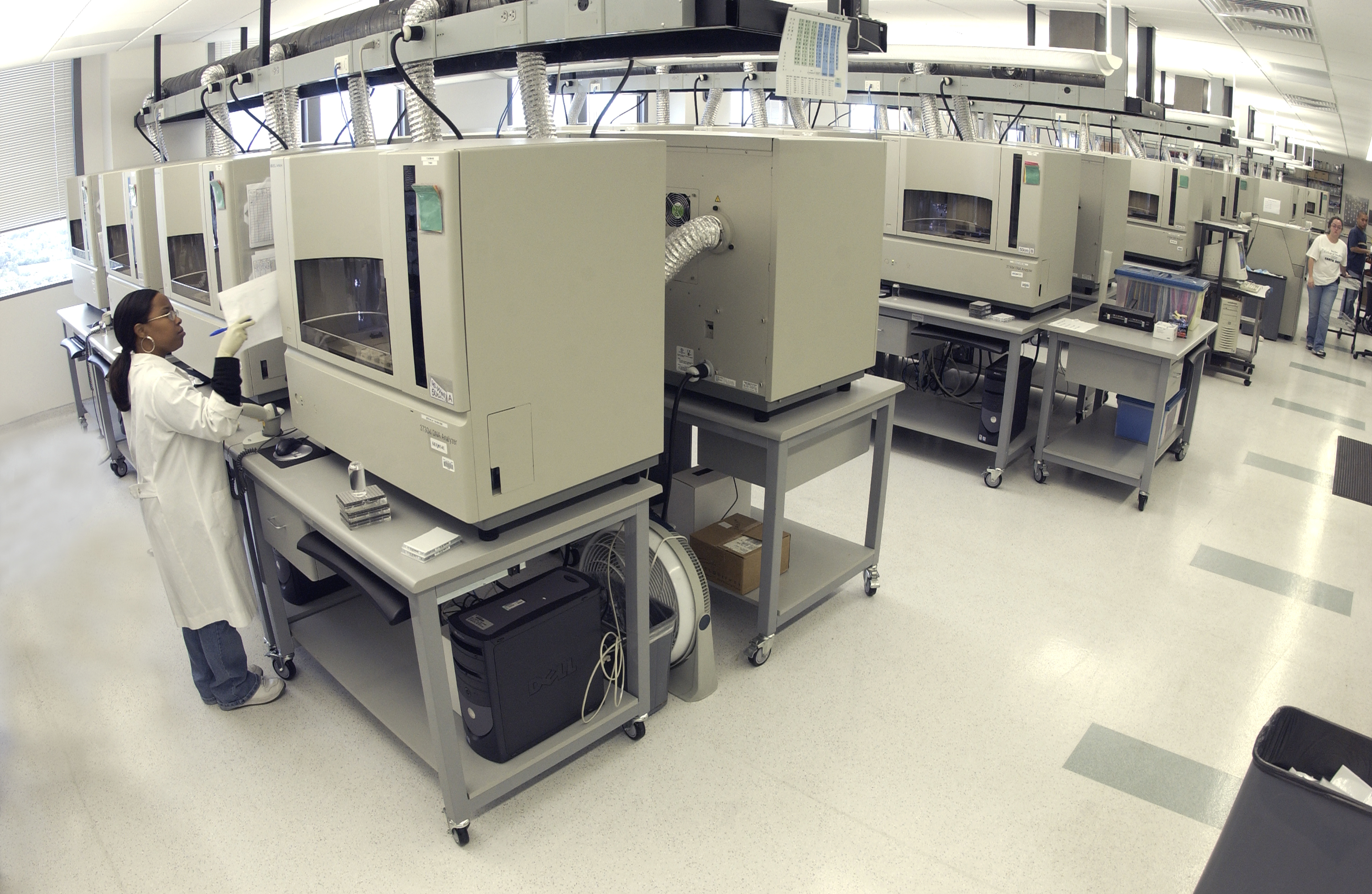
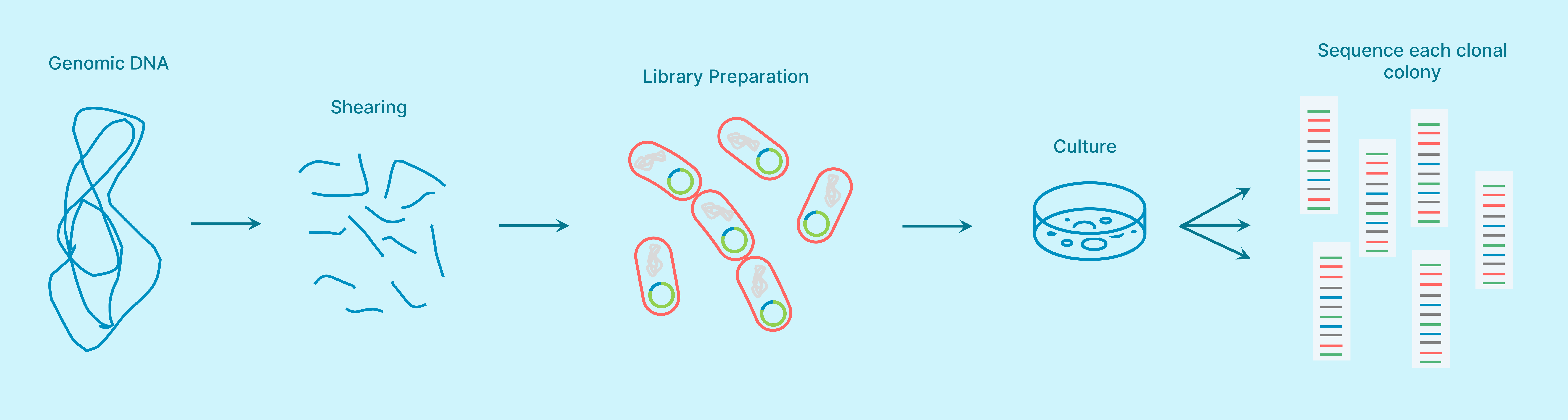
DNA Sequencing
A molecular technique to reveal the arrangement of nucleotides in polynucleotide chains...
First Generation
Wandering Spot Analysis (1973)
- Developed by Maxam & Gilbert
- Chemical Sequencing Method
- Chemical Cleavage Method
- Uses chemicals to degrade the sequence at particular nucleotides
- Radioactive Phosphorus (P32)
- Formic Acid (For depurinating Purines)
- Dimethyl Sulfate (For methylating Guanines)
- Hydrazine (For hydrolyzing Pyramidines)
- Hydrazine with NaCl (For removing Cytosine)
- Piperidine (For cleaving Phosphodiester backbone)
- Upto 500 nucleotides
- Extensive use of hazardous chemicals
- Difficult to develop kits
Sequence: ATGCAGCGTTA
| G |
AT ATGCA ATGCAGC ATGCAGCGTTA
|
|---|---|
| A+G |
AT ATGC ATGCA ATGCAGC ATGCAGCGTT
|
| C |
ATG ATGCAG
|
| C+T |
A ATG ATGCAG ATGCAGCG ATGCAGCGT
|

Chain Termination Sequencing (1977)
- Developed by Fredrick Sanger
- Sanger Sequencing
- Dideoxy Method
- Sequence-by-Synthesis (SBS)
- Uses ddNTPs to terminate the sequence
- dNTPs at higher concentration than the ddNTPs
- Approximately 1000 bp of sequence can be run in single reaction
- Automated Capillary Electrophoresis
- Major Provider: Applied Biosystems
- Output format: ab1
- Tools: BioEdit, Chromas, MEGA, etc.
Sequence: ATGCAGCGTTA
| A |
A ATGCA ATGCAGCGTTA
|
|---|---|
| T |
AT ATGCAGCGT ATGCAGCGTT
|
| G |
ATG ATGCAG ATGCAGCG
|
| C |
ATGC ATGCAGC
|
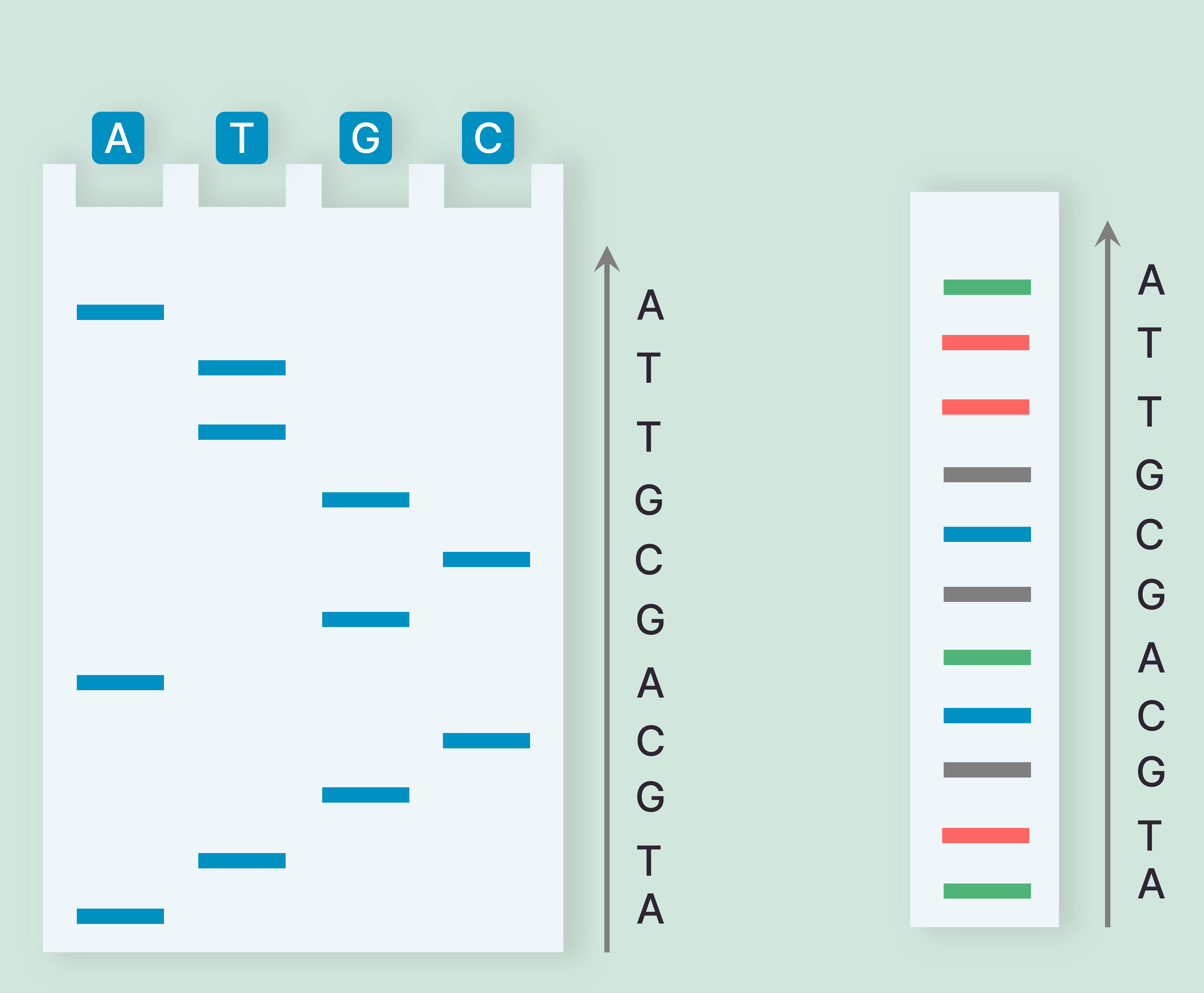
Whole Genome Sequencing with Sanger's Method